Hunting enzymes in the membrane: A new strategy to inhibit rhomboid proteases
IOCB Prague: Hunting enzymes in the membrane: A new strategy to inhibit rhomboid proteases
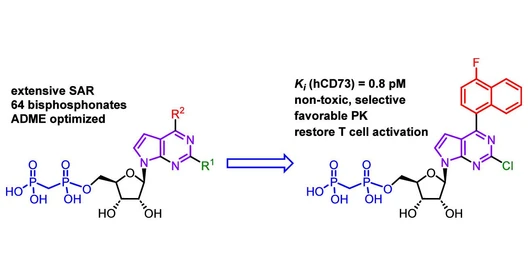
Rhomboid proteases are intriguing drug targets because they sit inside cell membranes of almost all organisms and control key biological processes linked to health and disease. Yet designing selective inhibitors has been notoriously difficult, as their active sites are similar and structural data are scarce. In addition, they work in the oily environment of the membrane, which complicates both experiments and drug design.
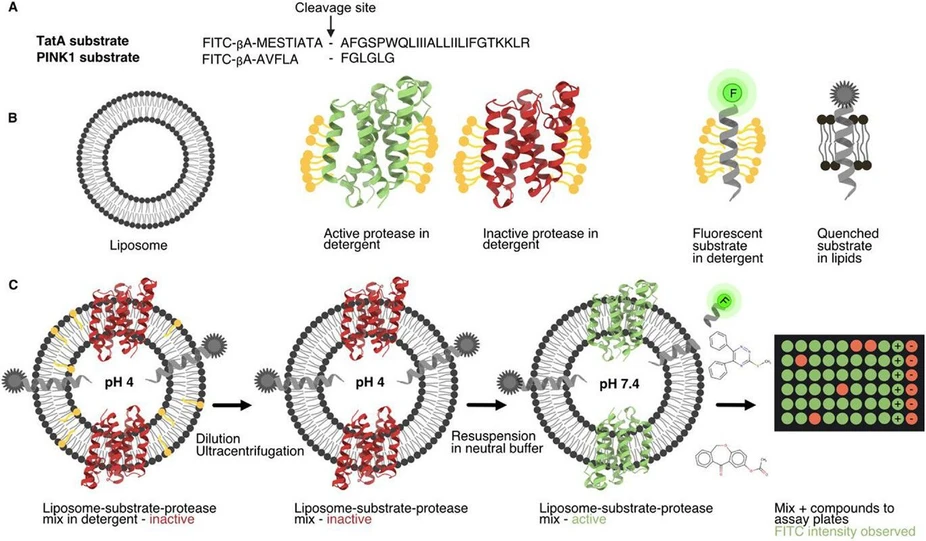
 IOCB Prague: Hunting enzymes in the membrane: A new strategy to inhibit rhomboid proteases: Assay scheme.
IOCB Prague: Hunting enzymes in the membrane: A new strategy to inhibit rhomboid proteases: Assay scheme.
The researchers led by Kvido Stříšovský from IOCB Prague and Adam Lange from Leibniz Forschungsinstitut für Molekulare Pharmakologie in Berlin built a high-throughput screening platform that mimics the natural membrane setting and uses a native-like substrate. This physiologically relevant setup led to the discovery of several chemically diverse, non-peptidic inhibitors that selectively block the bacterial rhomboid GlpG protease, which is a key pathogenic factor in uropathogenic, biofilm-forming bacteria.
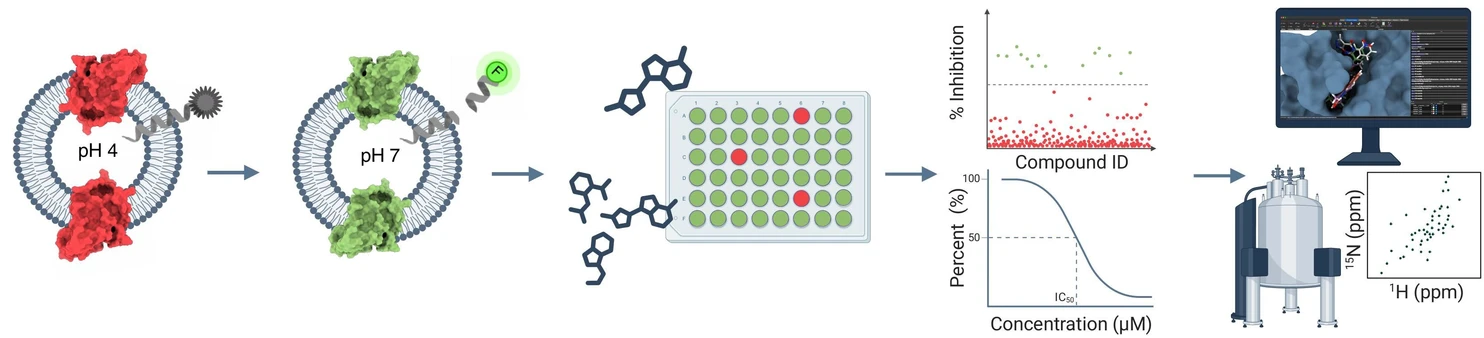 IOCB Prague: Hunting enzymes in the membrane: A new strategy to inhibit rhomboid proteases.
IOCB Prague: Hunting enzymes in the membrane: A new strategy to inhibit rhomboid proteases.
One of the active compounds showed particularly promising potency against both GlpG and the human mitochondrial rhomboid PARL, which regulates mitochondrial quality control and stress responses and has been implicated in the pathogenesis of Parkinson’s disease. Analogues of another hit compound revealed a new class of reversible covalent inhibitors that even function in living bacteria with permeabilized membranes. By combining docking studies, structural analysis of binding mode by solid-state NMR, and structure–activity analysis aided by computational modeling, the joint international team uncovered how these molecules bind and interact with critical catalytic features of the enzyme.
Beyond identifying new inhibitor scaffolds, the study provides a blueprint for targeting rhomboid proteases in disease contexts such as Parkinson’s disease, malaria, bacterial biofilms in the urinary tract and other infections, which is an important step toward future therapeutic applications.
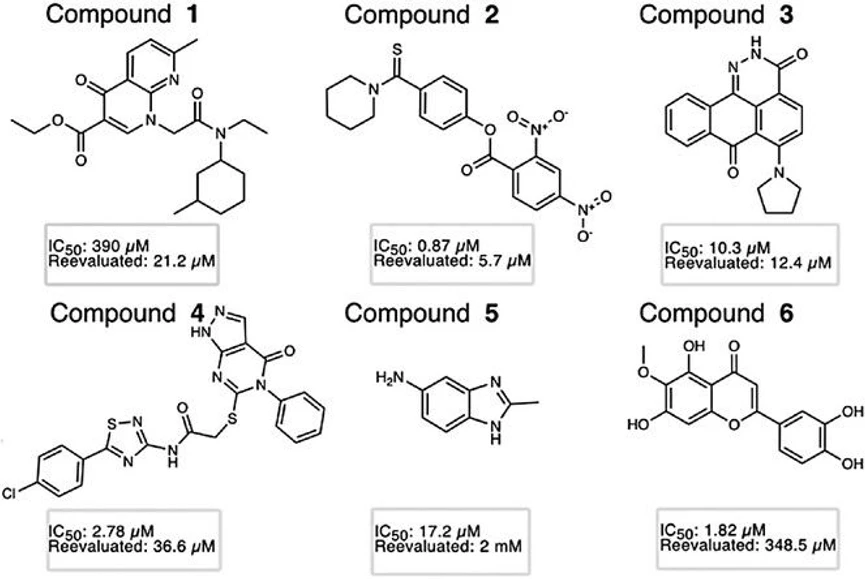 IOCB Prague: Hunting enzymes in the membrane: A new strategy to inhibit rhomboid proteases: Chemical structures of the 6 final compounds selected for further characterization with favorable inhibition profiles on FL GlpG.
IOCB Prague: Hunting enzymes in the membrane: A new strategy to inhibit rhomboid proteases: Chemical structures of the 6 final compounds selected for further characterization with favorable inhibition profiles on FL GlpG.
The Original article
Targeting the Membrane-Embedded Rhomboid Protease GlpG: A Multimodal Strategy for Inhibitor Discovery and Mechanistic Insight
Claudia Bohg, Yurii Dubanych, Spyridon Kosteletos, Taoran Xiao, Martin Neuenschwander, Tillmann Utesch, Michael Lisurek, Carl Öster, Andreas Oder, Carola Seyffarth, Kathrin Bach, Denise-Liù Gracias Leone, František Filandr, Marc Wegert, Sascha Lange, Henry Sawczyc, Jens Peter von Kries, Christian P. R. Hackenberger, Edgar Specker, Han Sun, Kvido Stříšovský, Adam Lange
Angew. Chem. Int. Ed. 2026, 65, e14067
licensed under CC-BY 4.0
Abstract
Rhomboid proteases, a class of intramembrane proteases characterized by a Ser-His catalytic dyad, have recently emerged as promising therapeutic targets. While inhibitors for soluble serine proteases have been extensively studied, the spectrum of potent rhomboid protease inhibitor chemotypes is limited to active-site targeted nucleophiles. To address this limitation, we conducted a high-throughput screen of over 68,000 compounds targeting the E. coli rhomboid protease GlpG, using a fluorescent liposome-based assay. A selection of 326 inhibitory compounds was evaluated in a subsequent IC50 screen against two variants of GlpG (core domain and full length), a soluble serine protease (chymotrypsin), as well as the human mitochondrial rhomboid PARL. Of these, the selective inhibitory effects of 2 compounds and their analogues on GlpG were confirmed through further biochemical and biophysical characterisation, molecular docking, and solid-state NMR spectroscopy. This study paves the way for developing small-molecule tool compounds and drug-like molecules targeting rhomboid proteases.